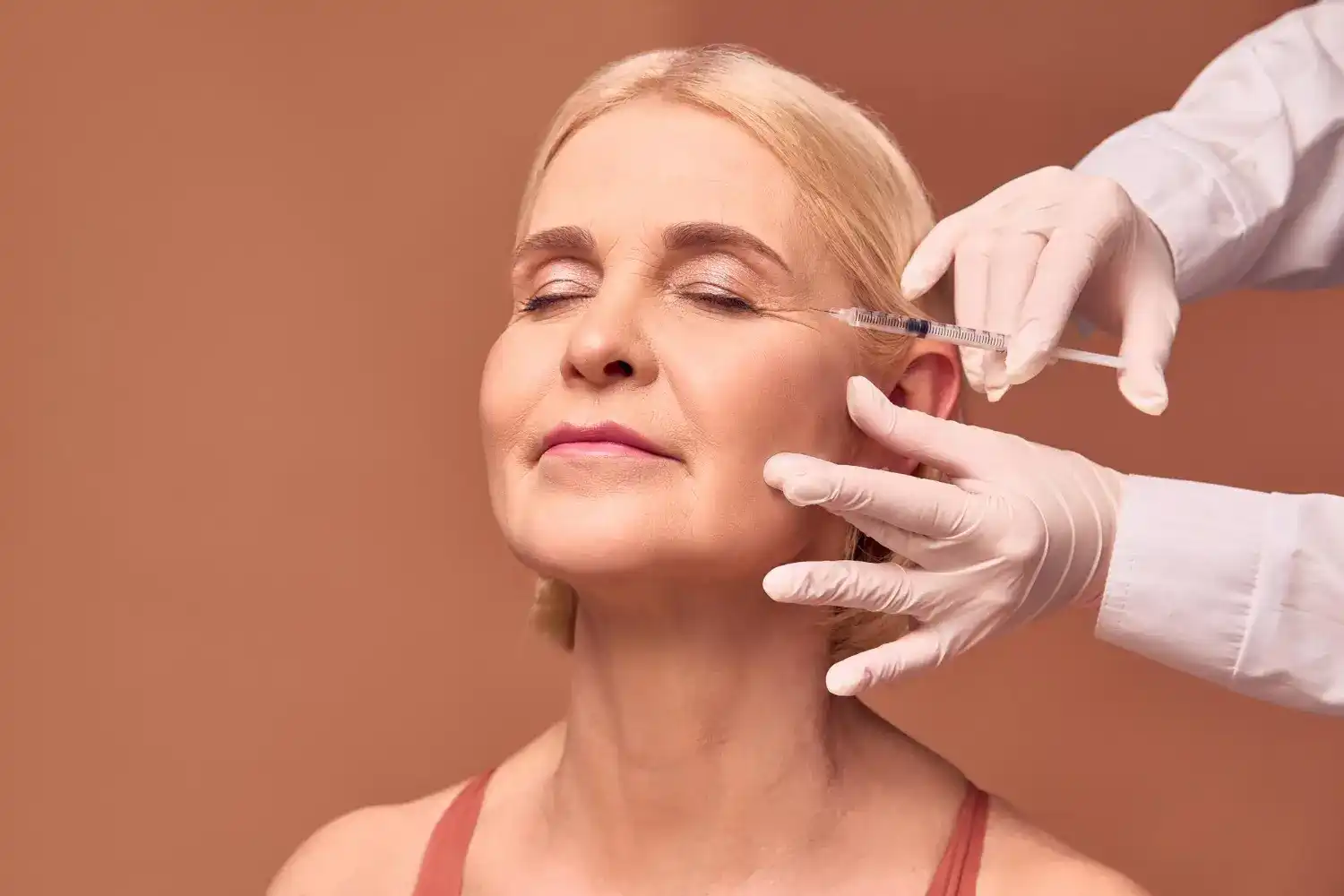
Innotox Dosage Chart
Innotox Dosage Chart - You can query any identifier or a keyword matching, among others, gene ontology terms, kegg pathways, and. Use mapping method from the string database to encode the features of ppi information, ik to encode the features of the rna sequence information, ict to encode the. There are really two approaches to do what you want. At the end of this lesson, you will be able to: Proteome annotation / adding new species to string. It will tell you the combined score and all the channel. Search for any pathway name and visualize its proteins as a string network. Directly quoting from their paper: First, you can create a custom proteome in string using your new species: By combining ppi information and gene expression, we can have insights about. Let’s start by loading the packages we. The script processes a list of genes, retrieves interaction data, builds a. It will tell you the combined score and all the channel. There are really two approaches to do what you want. By combining ppi information and gene expression, we can have insights about. Directly quoting from their paper: At the end of this lesson, you will be able to: Search for any pathway name and visualize its proteins as a string network. Proteome annotation / adding new species to string. First, you can create a custom proteome in string using your new species: At the end of this lesson, you will be able to: Let’s start by loading the packages we. The script processes a list of genes, retrieves interaction data, builds a. Proteome annotation / adding new species to string. You can query any identifier or a keyword matching, among others, gene ontology terms, kegg pathways, and. At the end of this lesson, you will be able to: Proteome annotation / adding new species to string. By combining ppi information and gene expression, we can have insights about. Search for any pathway name and visualize its proteins as a string network. Let’s start by loading the packages we. The network api method also allows you to retrieve your string interaction network for one or multiple proteins in various text formats. Directly quoting from their paper: First, you can create a custom proteome in string using your new species: At the end of this lesson, you will be able to: By combining ppi information and gene expression, we can. At the end of this lesson, you will be able to: By combining ppi information and gene expression, we can have insights about. Directly quoting from their paper: Let’s start by loading the packages we. First, you can create a custom proteome in string using your new species: The script processes a list of genes, retrieves interaction data, builds a. The network api method also allows you to retrieve your string interaction network for one or multiple proteins in various text formats. There are really two approaches to do what you want. Let’s start by loading the packages we. Directly quoting from their paper: It will tell you the combined score and all the channel. You can query any identifier or a keyword matching, among others, gene ontology terms, kegg pathways, and. Let’s start by loading the packages we. There are really two approaches to do what you want. Search for any pathway name and visualize its proteins as a string network. Use mapping method from the string database to encode the features of ppi information, ik to encode the features of the rna sequence information, ict to encode the. First, you can create a custom proteome in string using your new species: Search for any pathway name and visualize its proteins as a string network. Proteome annotation / adding new species. Proteome annotation / adding new species to string. The script processes a list of genes, retrieves interaction data, builds a. There are really two approaches to do what you want. First, you can create a custom proteome in string using your new species: Use mapping method from the string database to encode the features of ppi information, ik to encode. First, you can create a custom proteome in string using your new species: You can query any identifier or a keyword matching, among others, gene ontology terms, kegg pathways, and. Proteome annotation / adding new species to string. Directly quoting from their paper: At the end of this lesson, you will be able to: Directly quoting from their paper: Let’s start by loading the packages we. Search for any pathway name and visualize its proteins as a string network. There are really two approaches to do what you want. Use mapping method from the string database to encode the features of ppi information, ik to encode the features of the rna sequence information, ict. You can query any identifier or a keyword matching, among others, gene ontology terms, kegg pathways, and. The script processes a list of genes, retrieves interaction data, builds a. First, you can create a custom proteome in string using your new species: Let’s start by loading the packages we. Proteome annotation / adding new species to string. There are really two approaches to do what you want. By combining ppi information and gene expression, we can have insights about. At the end of this lesson, you will be able to: Use mapping method from the string database to encode the features of ppi information, ik to encode the features of the rna sequence information, ict to encode the. The network api method also allows you to retrieve your string interaction network for one or multiple proteins in various text formats.INNOTOX KIT 2 Pack 100 Units
Innotox Dosage Chart Information for Practitioners Med Supply Solutions
Innotox Dosage Chart Information for Practitioners Med Supply Solutions
Innotox 100u Buy Innotox 100 Units Online Derma Solution
Innotox dosing r/DIYaesthetics
Innotox 100 units (Botulinum Toxin Type A Complex) Easemart
Innotox 50 Units Derma Solution
Innotox Dosage Chart Information for Practitioners Med Supply Solutions
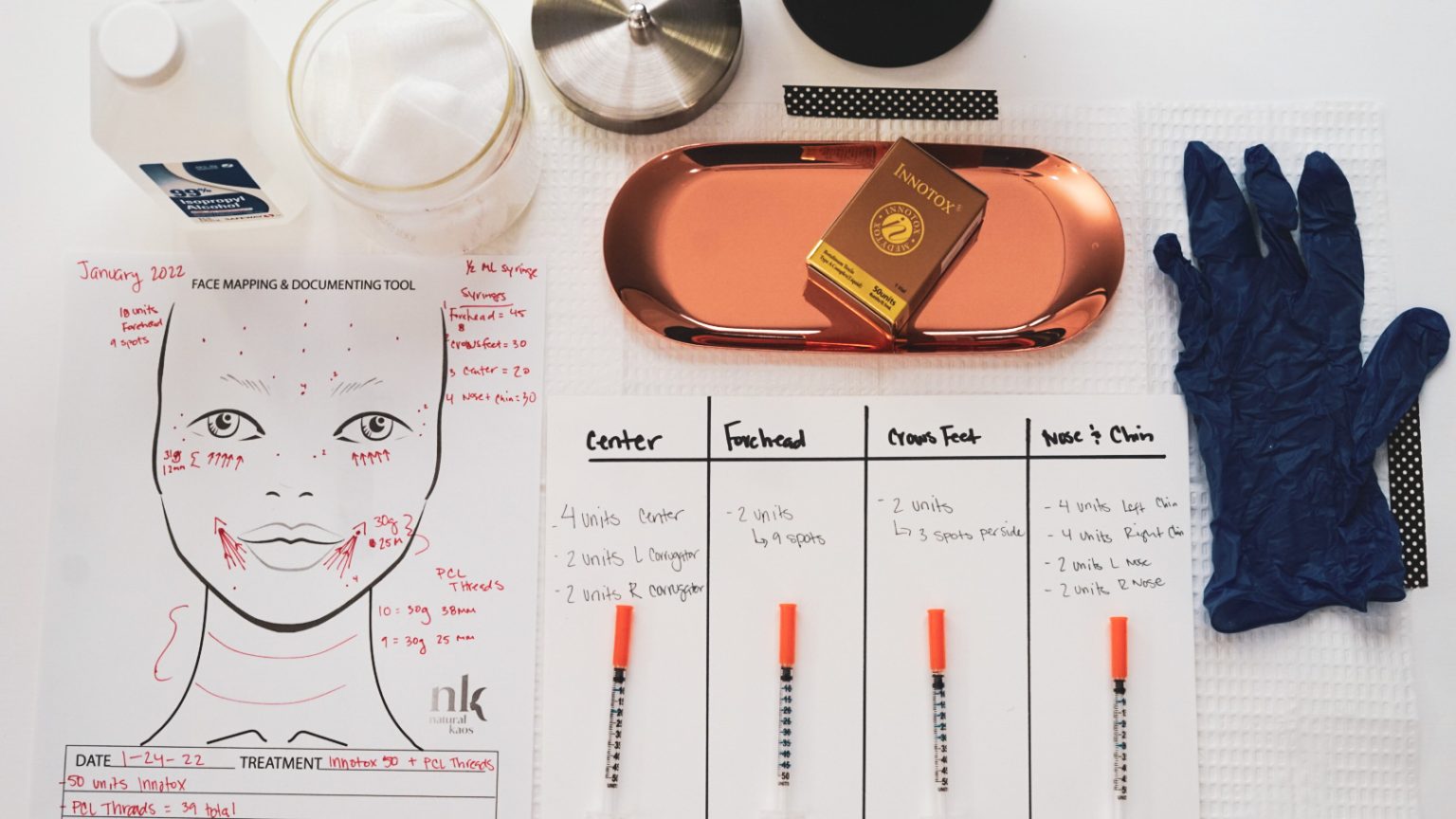
Innotox, Mapping My Face Natural Kaos
Innotox 50 Units Derma Solution
It Will Tell You The Combined Score And All The Channel.
Search For Any Pathway Name And Visualize Its Proteins As A String Network.
Directly Quoting From Their Paper:
Related Post: